
Contact
Email: eatoth@umd.edu
Call: (240) 314-6516
Eric Toth
Associate Research Professor
Toth Group
Contact
Email: eatoth@umd.edu
Call: (240) 314-6516
Education
- Postdoctoral Fellowship, University of California, Los Angeles (UCLA), and Harvard Medical School, 2000-2004
- Ph.D., Biochemistry, UCLA, 1999
- B.A., Biochemistry (Honors), University of Pennsylvania, 1992
Profile
Dr. Eric Toth applies biochemical and biophysical techniques, including X-ray crystallography, to accelerate the development of agents that modulate the function of a wide array of potential therapeutic targets. These efforts include the development of next-generation protein therapeutics, novel vaccines, and small molecule inhibitors of biologically important proteins.
CURRENT RESEARCH
Therapeutic Protein Design and Development
One area of research in Dr. Toth’s laboratory is the engineering of well-characterized proteins to develop new properties for use as therapeutics, sensors, or novel reagents. The development of a protein-based therapeutic directed against cancer-causing Ras protein is a proof-of-principle project currently underway. This project is a collaborative effort conducted by a team composed of Dr. Toth, Dr. John Orban (UMCP Department of Chemistry and Biochemistry) and Dr. Phil Bryan (Potomac Affinity Proteins), with contributions from Dr. Silvia Muro (IBBR) and the Ras Initiative (NCI-Frederick).
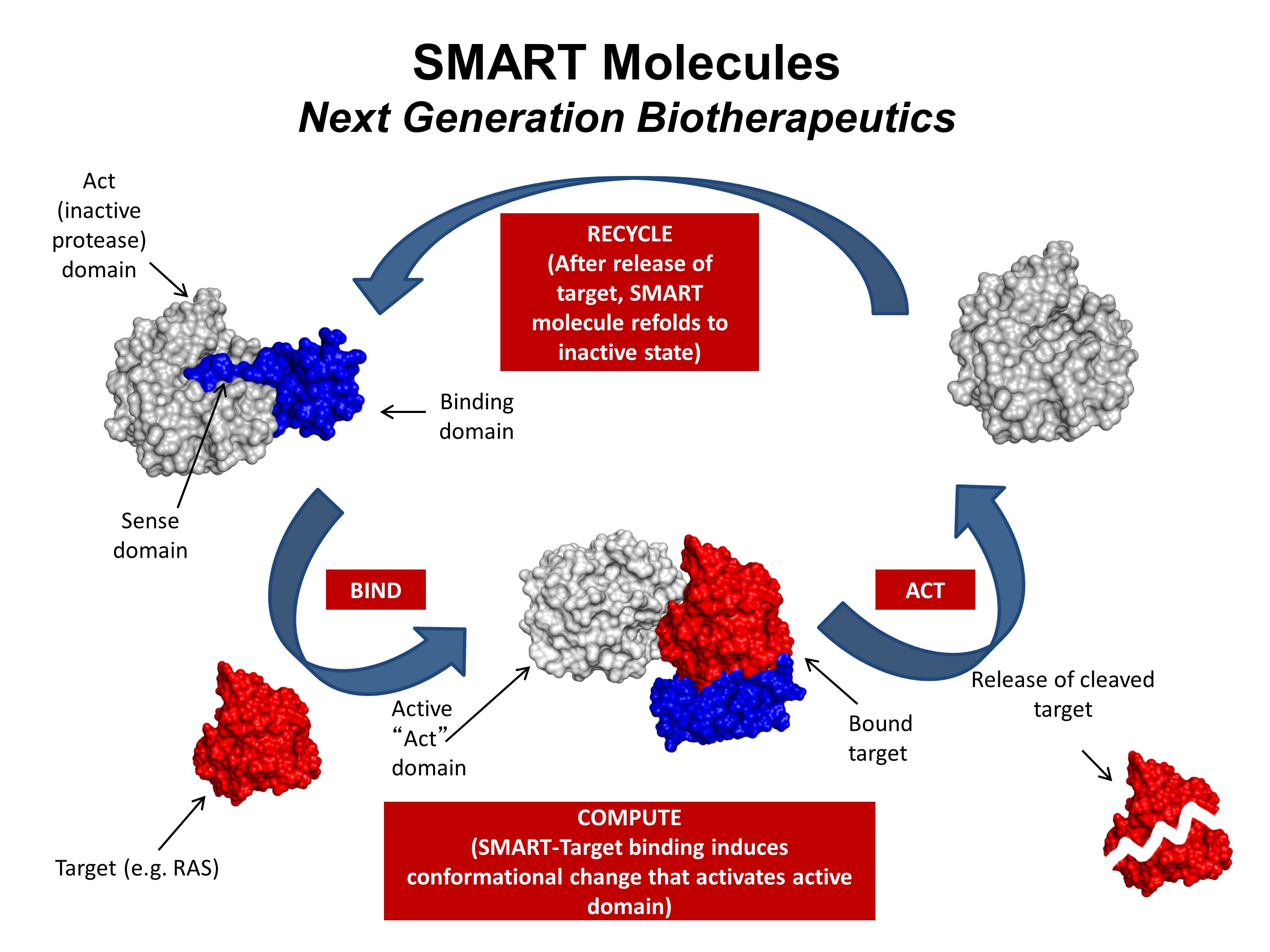
Mutations in one of the three RAS genes are involved in roughly a third of all human cancers, but therapeutic interventions have limited clinical efficacy. The goal of this project is to engineer a highly specific, highly regulated protease capable of destroying mutated human Ras proteins. The long-term objectives are to develop a “smart” therapeutic and to create a platform for future engineering of enzymatic machines to treat drug-resistant cancers and other diseases.
Hepatitis C vaccine
The Toth lab is part of a multi-institutional team, led by IBBR director Dr. Thomas Fuerst, which is developing a vaccine to prevent hepatitis C viral infection. Funded by a $6 million NIH grant, the effort takes a structure-based vaccine design approach to engineer vaccine candidates that elicit a broadly neutralizing immune response. Dr. Toth’s lab is developing novel methods for producing and isolating candidate vaccine antigens for biochemical and immunological testing.
Hepatitis C virus (HCV) is a major cause of severe liver disease and cancer with a global burden of nearly 185 million infected individuals. HCV is an RNA virus that mutates rapidly, making both treatment and vaccine development challenging for this pathogen. While HCV-specific antiviral agents provide effective therapy, a successful treatment does not prevent reinfection. In addition, the high cost of these drugs restricts access, particularly in developing nations where disease burden is greatest, further underscoring the need for a vaccine.
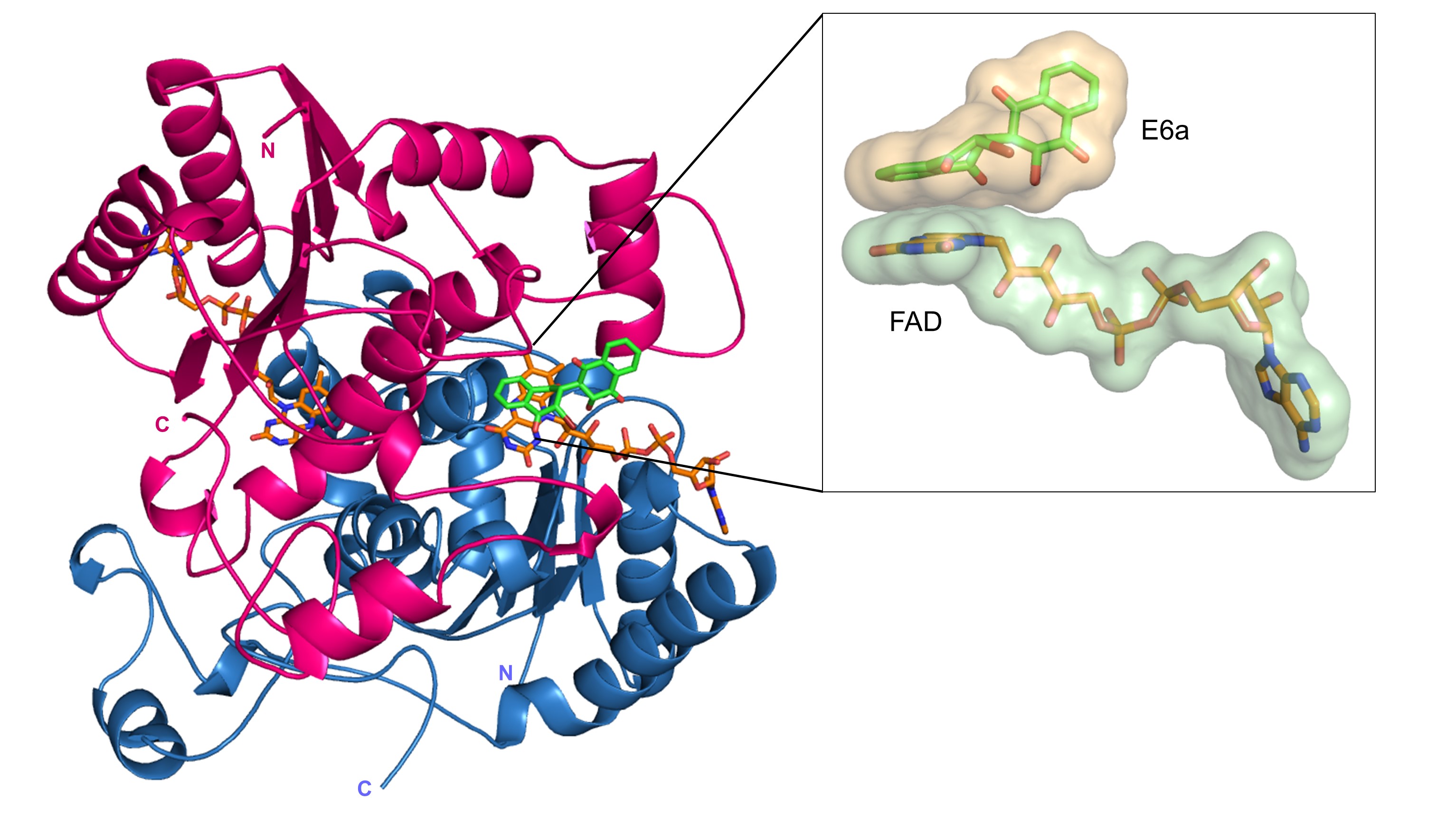
Structure-Based Drug Design
Dr. Toth has led efforts to determine crystal structures of several important drug targets in complex with novel lead compounds aimed at combating cancer and neurological diseases. These efforts include targeting malignant melanoma through disruption of the interaction of p53 with S100B, combating acute myeloid leukemia and other cancers with novel naphthoquinones that inhibit NQO1 (see inset), and targeting neurological disorders by developing inhibitors of the kynurenine pathway of tryptophan degradation.