Structural Genomics
Structural Genomics seeks to determine the three-dimensional structures of all protein families emerging from genome sequencing projects.
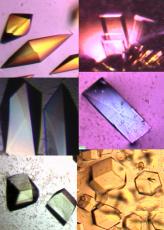 Our Structure to Function (S2F) project focused initially on bacterial proteins of unknown function. The goals were to infer function from structure when the target protein adopted a fold that was already available for proteins of known function, and to fill in the fold space in cases for which the structure revealed new fold. Over the course of these studies we found that 75% of the structures exhibited known folds, in which case we obtained clues about the general biochemical function. For 25% of the structures, we discovered new folds. When generic assays could be performed, such as phosphatase assay or DNA/RNA gel shift, the general biochemical function of the protein could be confirmed.
Our Structure to Function (S2F) project focused initially on bacterial proteins of unknown function. The goals were to infer function from structure when the target protein adopted a fold that was already available for proteins of known function, and to fill in the fold space in cases for which the structure revealed new fold. Over the course of these studies we found that 75% of the structures exhibited known folds, in which case we obtained clues about the general biochemical function. For 25% of the structures, we discovered new folds. When generic assays could be performed, such as phosphatase assay or DNA/RNA gel shift, the general biochemical function of the protein could be confirmed.
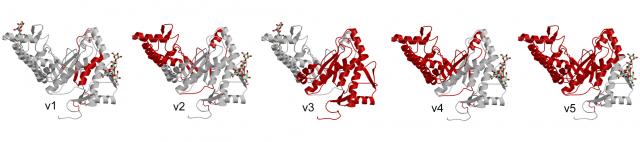
We now study the structural consequences of alternative splicing. Alternative splicing occurs in high eukaryotes and provides the organism with the capacity to expand the scope of functions beyond the number of genes encoded by the DNA. Our goal is to provide better understanding of the impact of the structure on function.