Contact
Email: arlin@umd.edu
Call: (240) 314-6208
Arlin Stoltzfus
Research Biologist
Stoltzfus Group
Contact
Email: arlin@umd.edu
Call: (240) 314-6208
Education
- Postdoctoral Fellow, Dalhousie University, 1999
- Ph.D. Biology, University of Iowa, 1991
- B.A., English, Grinnell College, 1985
Profile
Dr. Stoltzfus's research addresses issues in molecular evolution, bioinformatics, and evolutionary theory that are amenable to computer-based approaches. The group explores models and tests hypotheses, develops software, and participates in community efforts to improve interoperability. A major ongoing interest is understanding the role of mutation in evolution, an area where the group has developed novel theory and presented novel results on the foundational issue of how biases in mutation influence the course of evolution. Other topics of interest include the evolution of introns, Bayesian methods for fossil calibration of phylogenies, models of constructive neutral evolution, and the history and philosophy of evolutionary biology.
CURRENT RESEARCH
Mutation-biased evolution
Drs. Yampolsky and Stoltzfus (2001) showed that mutational and developmental biases in the introduction of variation can influence evolution, contrary to classical arguments that selection controls the course of evolution because mutation "pressure" is weak. Recently, a wealth of information on cases of parallel molecular adaptation has made it possible to validate the distinctive prediction that mutation biases can be effectual even during adaptive evolution (Stoltzfus and McCandlish, 2017; Storz, et al., in press).
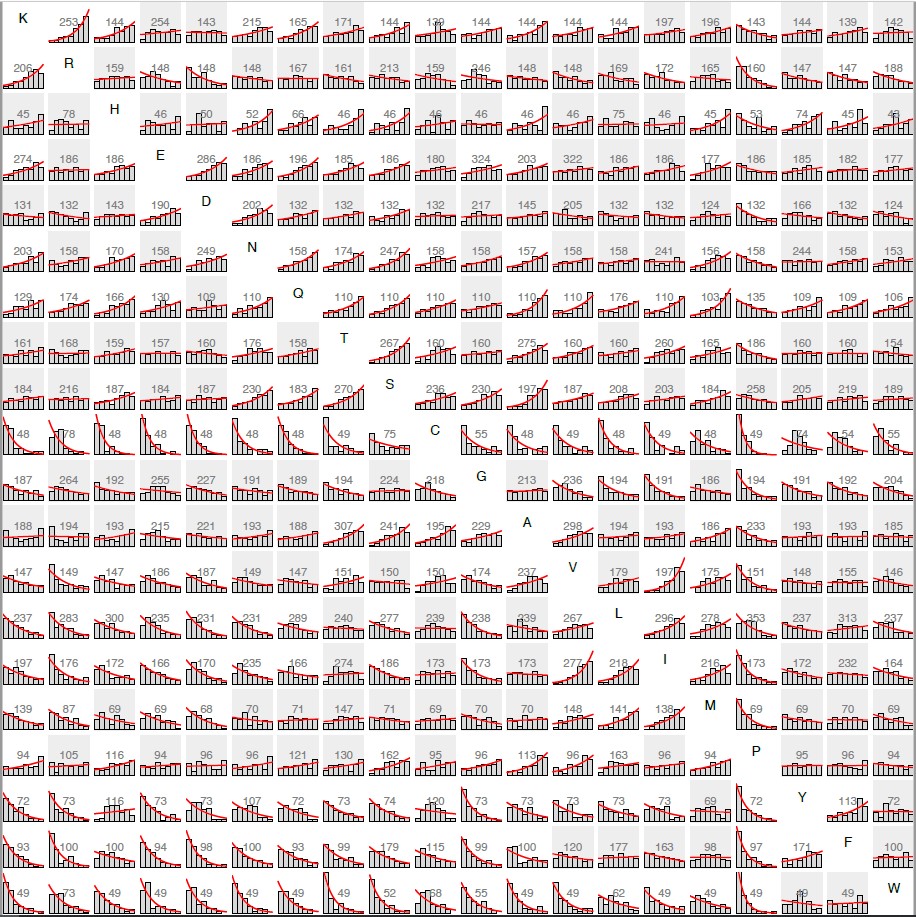
Meta-analysis of high-throughput fitness data
Drs. Yampolsky and Stoltzfus (2005) were the first to use experimental data to develop an empirical model of amino acid exchangeability. Drs. Stoltzfus and Norris (2015) used fitness data on over 1,000 protein mutants to show that, contrary to conventional wisdom in molecular evolution, transition mutations that change amino acids in proteins are not meaningfully more conservative than transversions. Ongoing work on amino acid exchangeability involves dozens of high-throughput, "deep mutational scanning" studies covering over 105 mutants (McCandlish and Stoltzfus, in preparation).
Disseminating the Tree of Life
With collaborators at the University of Tennessee, Knoxville, New Mexico State University, and the University of Illinois Urbana-Champaign, the Stoltzfus group is building a "Phylotastic" system to extract species trees from existing resources on the fly. The Phylotastic web portal (http://portal.phylotastic.org) provides several user-friendly workflows for obtaining species trees.