Pierce Lab
Links
We develop and apply algorithms to model and design protein-protein interactions, including ZDOCK, ZRANK, Rosetta, and TCRFlexDock. With a major focus on immune recognition and therapeutics, we are particularly interested in T cell receptors, antibodies, and vaccine design.
For tools, resources, and updated lab information, please see our lab web site here:
http://piercelab.ibbr.umd.edu.
Additional information on the lab's research can be found below (not updated regularly).
Vaccine Design
Advances in computational design algorithms, coupled with experimentally determined data on the structural basis of immune recognition and pathogenesis of viruses, viral sequence and immune repertoire sequencing, are ushering in a new era of vaccine design to combat emerging and variable viruses.
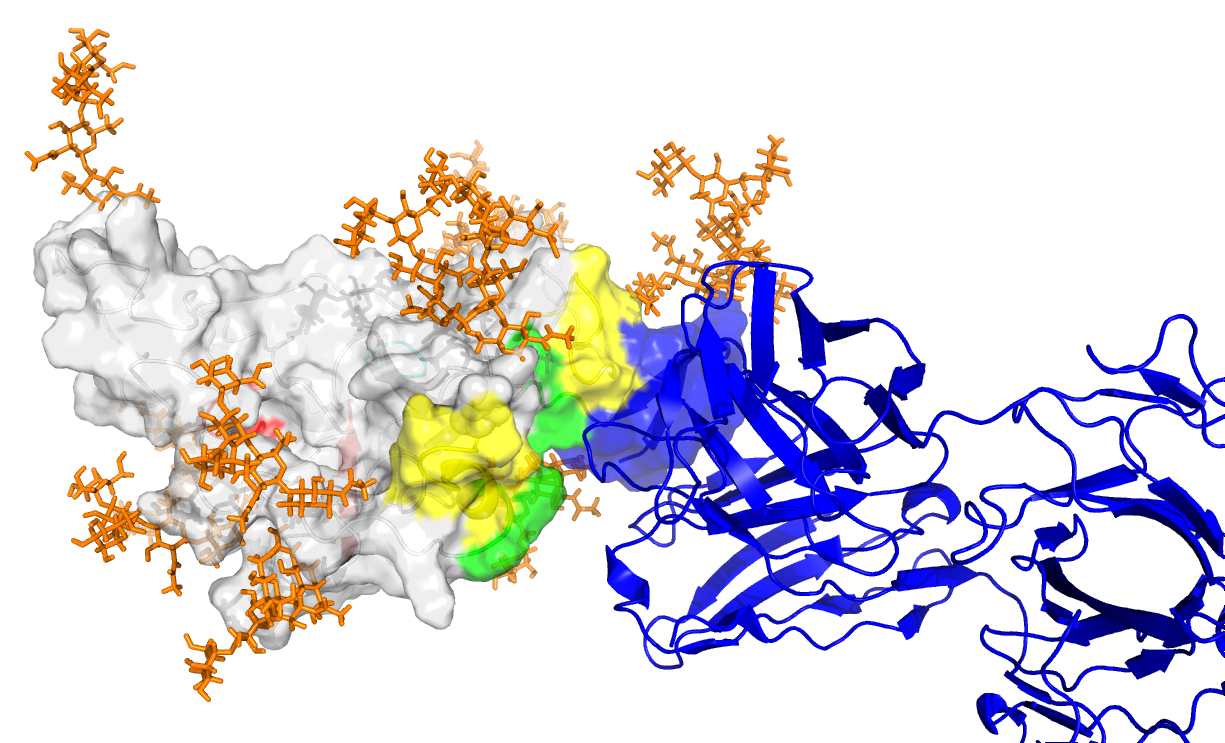
We have been studying hepatitis C virus (HCV) and design of vaccines to elicit antibodies to target conserved and functionally critical portions of the viral envelope. Through collaborations with leading HCV researchers, including MassBiologics who recently had their HCV1 mAb in phase 2 clinical trials, we are engineering and testing a number of novel vaccine candidates to effectively target this virus.
TCRs
T cell receptors (TCRs) are molecules of the adaptive immune system, similar to antibodies, that specifically bind peptide antigens loaded onto MHC molecules on the cell surface, and are of high interest as soluble and cell-based immunotherapeutics.
TCR engineering for high affinity and specificity
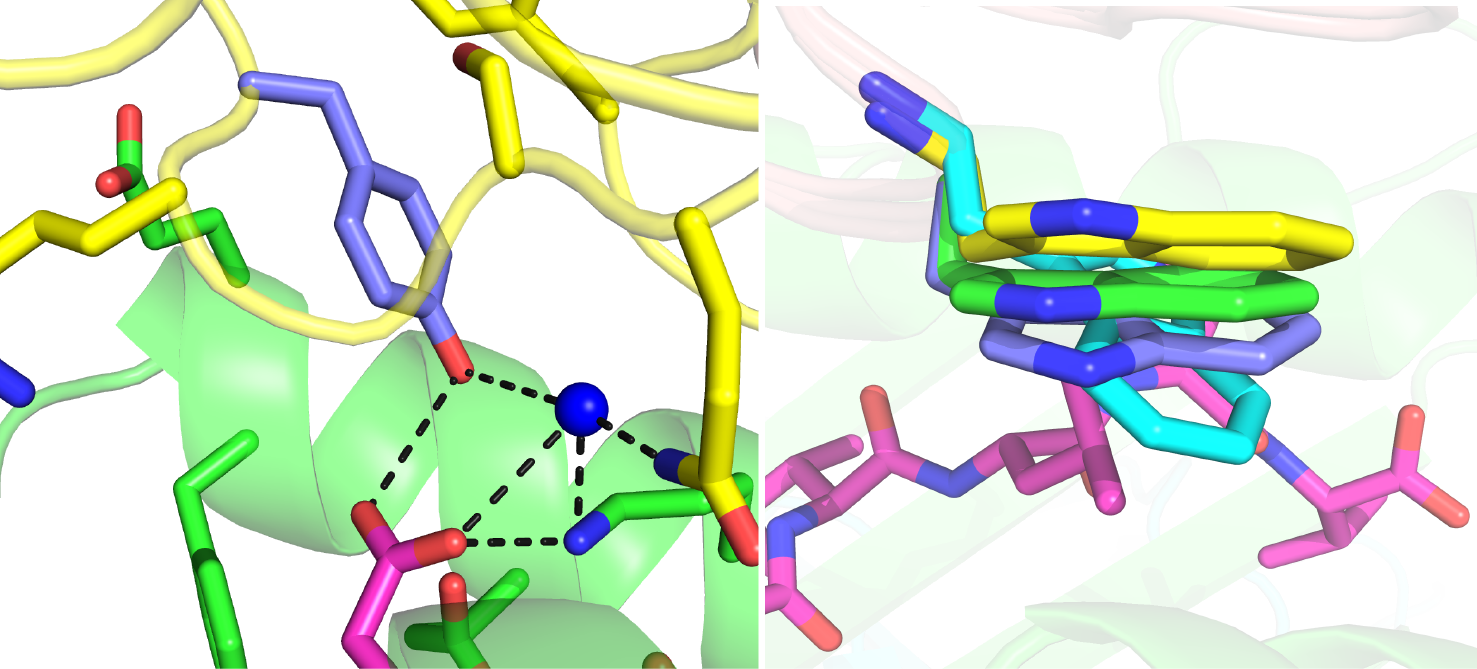
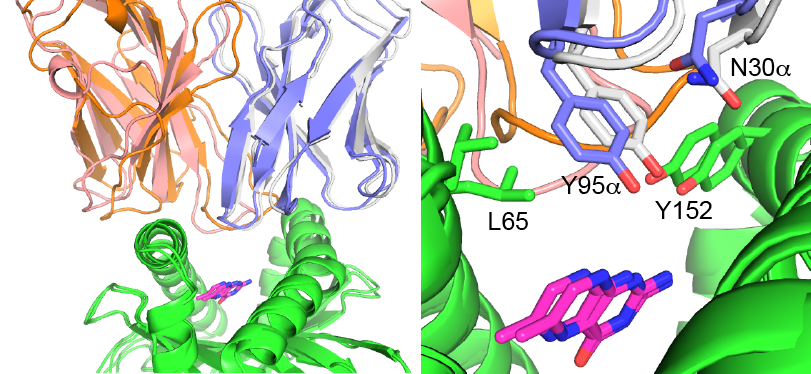
To enhance their targeting and therapeutic capabilities, we have engineered several TCRs to improve their pMHC affinities by up to 100-400-fold. In collaboration with the Baker laboratory at Notre Dame, the structure of our high affinity TCR engineered to better target a tumor antigen was solved. We are working to design the affinity of other TCRs and to alter and redirect antigen specificities.
Modeling TCR recognition
We are also making major strides toward a predictive understanding of TCR-pMHC recognition, having developed the first TCR-pMHC docking algorithm, TCRFlexDock. This was validated on a benchmark of 20 unbound TCR-pMHC test cases, and was also recently applied to predict TCR recognition of MHC-like molecules CD1 and MR1 bound to various lipid and small molecule antigens.
Antibodies
Our interest in antibody recognition is two-fold:
- Design broadly neutralizing and therapeutic antibodies
- Predict the structure and affinity of antibody-antigen interactions
Using computational protein design and docking algorithms, such as Rosetta, ZAFFI, and ZDOCK we are exploring ways to improve antibody targeting of viruses and to better predict uncharacterized interactions.
