IBBR Researchers Release New Database to Support Immunology Research
Wed, Jul 10, 2019
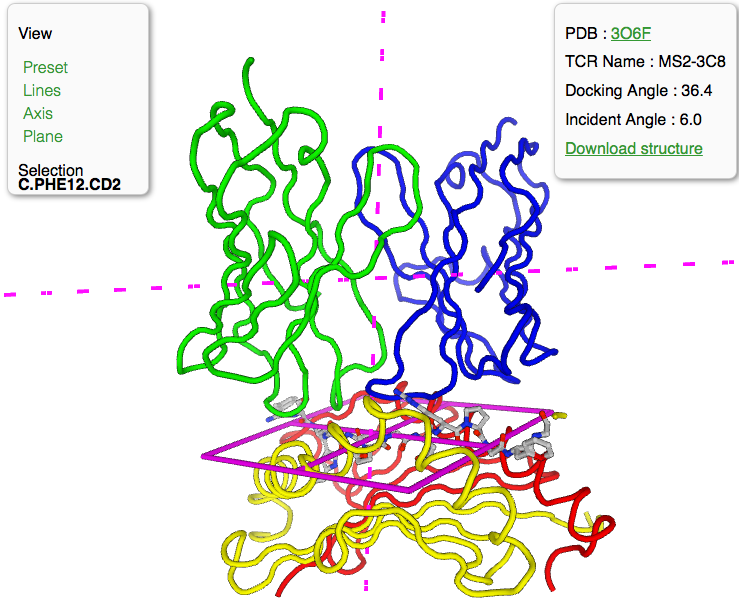
July 11, 2019 - IBBR Fellow Dr. Brian Pierce (Assistant Professor, UMCP Department of Cell Biology and Molecular Genetics) and postdoctoral associate Dr. Ragul Gowthaman recently released a new tool for researchers who study immune system proteins. The TCR3d database contains all known T cell receptor (TCR) structures and is updated weekly.
TCRs recognize proteins from infectious agents and signal the immune system to respond. When TCRs mistakenly recognize the body’s own proteins, autoimmune disorders can result. These highly specific immune signaling proteins represent a growing class of therapeutics, and a number of clinical trials of TCR-based therapies for cancer are in progress.
The TCR3d database particularly focuses on structures of TCRs in complex with various targets. Its simple and accessible interface will allow researchers to search for patterns to better understand TCR binding and facilitate the development of novel therapeutics and vaccines.
“We are thrilled to offer this database to the community, to permit researchers to gain improved insights into TCR recognition, and to enable better understanding of the vast number of structurally uncharacterized TCRs,” says Pierce. “We thank our collaborators who provided useful input on the database, the IBBR IT staff, and the many structural biologists whose work resulted in these fascinating structures.”
This tool complements TCRmodel, a web server for modeling T cell receptor structures based on protein sequence, which the team published and released in 2018.
This work was supported by the National Institutes of Health (Award R01-GM126299 to Brian Pierce). The article describing the database appears in the journal Bioinformatics: https://doi.org/10.1093/bioinformatics/btz517
Initial funding for Pierce’s research was provided through a seed grant from the University of Maryland Strategic Partnership: MPowering the State, a program designed to leverage the strengths and missions of the University of Maryland College Park and the University of Maryland Baltimore.
-----
Inquiries: communications@ibbr.umd.edu
