Development of the Glycoengineered CHO Cell Panel
Mammalian cells have many glycosyltransferases synthesizing the glycan structures on secreted proteins, and a few dozen enzymes that are responsible for adding the abundant N-linked glycans on recombinant proteins. CHO cells express most glycosyltransferases involved in mammalian N-linked glycosylation, with the key enzymes for making complex and hybrid N-glycans. However, the enzymes synthesize the glycans through complex pathways. Thus, for a given glycoprotein, often a diverse set glycoforms are made, with subtle differences in the glycan structures decorating the protein. Unfortunately, this heterogeneity leads to two problems. First, when glycosylation effects function, heterogeneity dilutes the efficacy of the protein product and this can also lead to safety issues. Second, heterogeneity results in more glycans whose abundance must be controlled, if possible, during scale-up and manufacturing once the product is approved. Thus, there is a need to identify which glycoforms possess the most desirable properties and then to develop cell lines producing the optimal glycoform as homogeneously as possible.
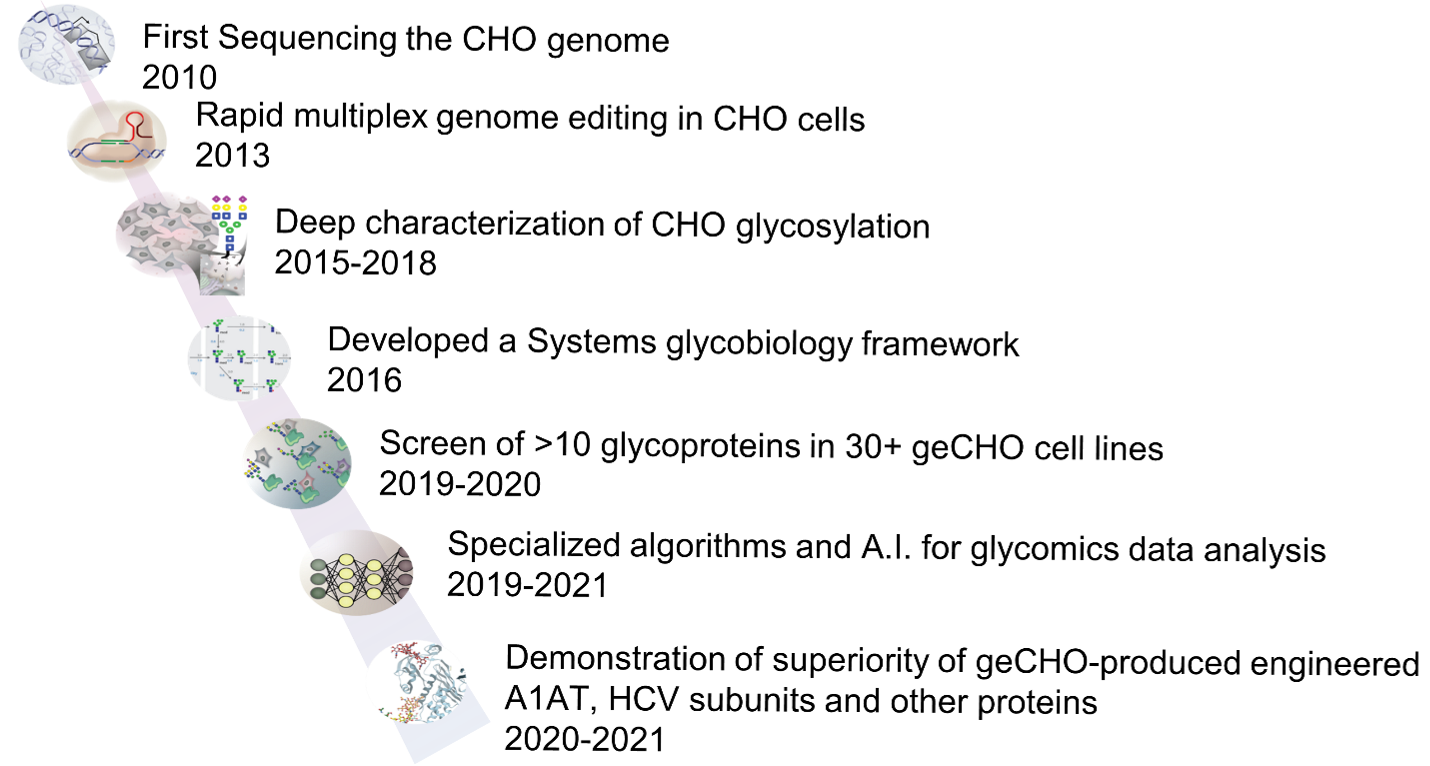
To achieve these goals, we deeply characterized glycosylation in CHO cells (Fig. 1). First, we sequenced and annotated the CHO-K1 and Chinese hamster genomes 1,2,3 and quantified glycosyltransferase mRNA expression. Furthermore, we and others linked each glycosyltransferase to their functions by knocking out (or knocking in) each gene individually and in combination 4,5,6,7 , and then quantified the glycome and transcriptome. Further using specialized systems biology modeling techniques, we were able to assign their role in glycosylation, including primary enzyme specificities and off-target effects. Using this information, over the course of 10 years we developed a panel of 30 glycoengineered CHO cells (geCHO) each expressing a unique human glycoform. These cells were derived from CHO-S cells (Thermo Fisher), which can grow to industrial-level cell densities in suspension on serum-free media. We use these cells to produce diverse, but more homogeneous glycoforms of various recombinant proteins of interest.
A decade of pioneering systems glycoengineering in CHO
References
- 1. Xu X, Nagarajan H, Lewis NE, Pan S, Cai Z, Liu X, Chen W, Xie M, Wang W, Hammond S, Andersen MR, Neff N, Passarelli B, Koh W, Fan HC, Wang J, Gui Y, Lee KH, Betenbaugh MJ, Quake SR, Famili I, Palsson BO, Wang J. The genomic sequence of the Chinese hamster ovary (CHO)-K1 cell line. Nat Biotechnol. 2011 Jul 31;29(8):735-41. doi: 10.1038/nbt.1932. PMID: 21804562 . PMCID: PMC3164356 .
- 2. Lewis NE, Liu X, Li Y, Nagarajan H, Yerganian G, O'Brien E, Bordbar A, Roth AM, Rosenbloom J, Bian C, Xie M, Chen W, Li N, Baycin-Hizal D, Latif H, Forster J, Betenbaugh MJ, Famili I, Xu X, Wang J, Palsson BO. Genomic landscapes of Chinese hamster ovary cell lines as revealed by the Cricetulus griseus draft genome. Nat Biotechnol. 2013 Aug;31(8):759-65. doi: 10.1038/nbt.2624. Epub 2013 Jul 21. PMID: 23873082 .
- 3. Rupp O, MacDonald ML, Li S, Dhiman H, Polson S, Griep S, Heffner K, Hernandez I, Brinkrolf K, Jadhav V, Samoudi M, Hao H, Kingham B, Goesmann A, Betenbaugh MJ, Lewis NE, Borth N, Lee KH. A reference genome of the Chinese hamster based on a hybrid assembly strategy. Biotechnol Bioeng. 2018 Aug;115(8):2087-2100. doi: 10.1002/bit.26722. Epub 2018 May 29. PMID: 29704459 . PMCID: PMC6045439 .
- 4. Yang Z, Wang S, Halim A, Schulz MA, Frodin M, Rahman SH, Vester-Christensen MB, Behrens C, Kristensen C, Vakhrushev SY, Bennett EP, Wandall HH, Clausen H. Engineered CHO cells for production of diverse, homogeneous glycoproteins. Biotechnol Bioeng. 2015 Aug;33(8):842-4. doi: 10.1038/nbt.3280. Epub 2015 Jul 20. PMID: 26192319 .
- 5. Spahn PN, Hansen AH, Kol S, Voldborg BG, Lewis NE. Predictive glycoengineering of biosimilars using a Markov chain glycosylation model. Biotechnol J. 2017 Feb;12(2):10.1002/biot.201600489. doi: 10.1002/biot.201600489. Epub 2016 Dec 28. PMID: 27860290 . PMCID: PMC5293603.
- 6. Liang C, Chiang AWT, Lewis NE. GlycoMME, a Markov modeling platform for studying N-glycosylation biosynthesis from glycomics data. STAR Protoc. 2023 Apr 21;4(2):102244. doi: 10.1016/j.xpro.2023.102244. Epub ahead of print. PMID: 37086409 . PMCID: PMC10160804 .
- 7. Liang C, Chiang AWT, Hansen AH, Arnsdorf J, Schoffelen S, Sorrentino JT, Kellman BP, Bao B, Voldborg BG, Lewis NE. A Markov model of glycosylation elucidates isozyme specificity and glycosyltransferase interactions for glycoengineering. Curr Res Biotechnol. 2020 Nov;2:22-36. doi: 10.1016/j.crbiot.2020.01.001. Epub 2020 Jan 23. PMID: 32285041 . PMCID: PMC7153765.