CGAB Infrastructure
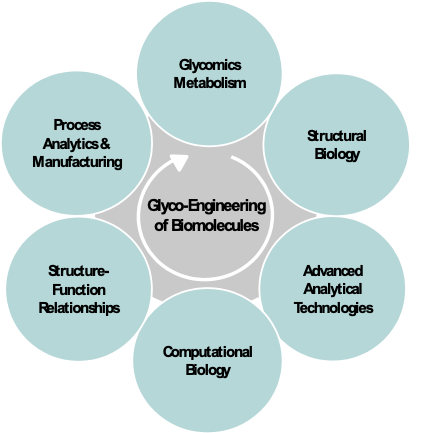
CGAB has assembled an outstanding core group of interdisciplinary investigators that have worked together over a decade spanning the disciplines of computational biology, understanding the fundamental pathways of glycobiology metabolism, control, and secretion, CHO cell line engineering, structural biology, protein production and purification, advanced protein analytical testing and characterization, molecular dynamics of structure-function relationships, bioproduction and process analytics, and quality control of biologics (Fig. 2). The team works in an industry-like, synergistic partnership to address critical research and development questions beyond the ability of any one lab. The group’s core science has focused on biotherapeutics and vaccines in which glycosylation and secretion play critical roles.
Production of a single recombinant glycoprotein can be laborious to deploy. However, the individual steps are highly automatable, including the steps for gene assembly, transfection, cell cultivation, and protein harvesting and production. To develop an automated workflow for cell line development and protein production, we established strong experimental and automation capabilities for cell and molecular biology. This includes our validated Opentrons
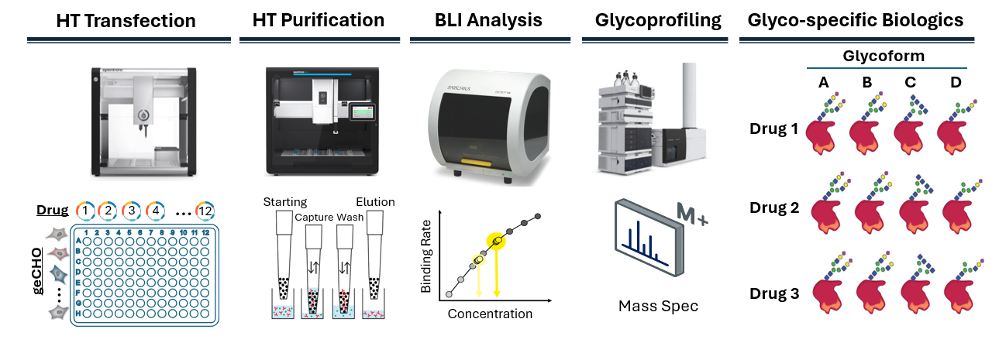
OT-2/Flex automation scripts for suspension-grown CHO-based production, which produces proteins in 96-well format (Fig. 3). This fully-automated, two-day workflow is published in the Opentrons Protocol Library, shows we can translate complex cell culture and glycoprotein expression workflows into robust, automation at scale.
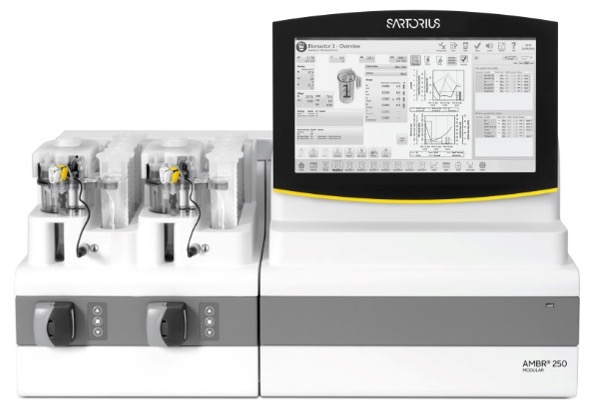

The CGAB partnership maintains world-class instrumentation and facilities to support development and advancement of bioscience measurements. At IBBR, these capabilities are maintained and operated to facilitate shared access to all IBBR researchers, through an organizational structure called the IBBR Commons. As part of that infrastructure, CGAB researchers will have access to testbeds for biomanufacturing research which are located on site at IBBR as part of the IBBR Commons NIST Biomanufacturing Resource Hub. This resource provides collaborators and stakeholders access to biopharmaceutical reference materials, measurement science data, and technology innovations occurring at the National Institute of Standards and Technology and IBBR. In particular, researchers have access to a fully-equipped cell culture facility for production of recombinant proteins in CHO cells. This includes an Ambr 250 bioreactor system (Fig. 4) that will allow preliminary process optimization and analysis at smaller scales using parallel cultures. This facilitates initial optimization of growth conditions or parallel production of a given glycoprotein in up to 24 geCHO cell lines for preliminary biochemical and glycan analysis. Available tangential flow filtration (TFF) equipment will speed up processing on both small and large scales. This facility is a collaborative effort by NIST and UMD and enables users to optimize conditions for CHO cell production of antibodies, biotherapeutics, and vaccines. IBBR also contains state-of-the art purification and characterization equipment, including two Akta Avant 150 chromatography systems for protein purification using industry-standard equipment. Use of the Avant allows for seamless portability of SOPs for vaccine production once the project reaches the initial preclinical development stage. Purified samples can then have an assessment of size and the distribution of species in solution via SEC-MALS using a Vanquish Flex UHPLC system coupled to Dawn HELEOS-II MALS and Optilab T-rEX refractive index detectors. Thermal stability can be assessed using a Nano DSC. Where applicable, users can measure biomolecular interactions via biolayer interferometry using an Octet RED96, surface plasmon resonance using a Bioacore T100, or isothermal calorimetry using a Microcal VP-ITC. Glycan content can be assessed either at IBBR using NIST resources or at UGA and the CCRC as described below.
The UGA Bioexpression and Fermentation Facility (BFF) is a cutting-edge facility located in the Department of Biochemistry and Molecular Biology at the University of Georgia. It offers state-of-the-art equipment and extensive expertise in protein production for both academic and industry clients. The BFF’s goal is to lead the way in research, development, and manufacturing, as well as to cultivate the next generation of scientists.

The Quality Control (QC) group has developed the noninvasive analytical technology wNMR (water nuclear magnetic relaxometry, Fig. 5) which can quantitatively and noninvasively analyze liquid drug products, including solutions, emulsions and suspensions. The wNMR platform has been applied to multiple marketed drug products, such as therapeutic proteins (e.g., Lantus and Neupogen), monoclonal antibody (e.g., Dupixent), vaccines (e.g., Daptacel) and small molecule drugs (e.g., Diprivan). The QC group also works with industry to characterize mAbs, vaccines and gene therapy products under development and has received support from the FDA, NIIMBL, NIST and TEDCO. The advantage of wNMR as an analytical technique is that it can analyze intact liquid drug products without opening the primary containers (vials, syringes, pens, bottles, etc.), and without any sample preparation and reagent. wNMR uses benchtop NMR instruments for analysis Fig. 5) in which a drug vial to be lowered into a benchtop NMR for analysis.
Additional QC analysis can entail detection of aggregates using a Zetasizer Nano ZS dynamic light scattering instrument. Confirmation of the identity of proteins produced in the Consortium and monitoring unwanted modifications such as oxidation or deamidation can be accomplished via the mass spectrometry multi-attribute method, which entails peptide mapping and analysis and an LTQ Orbitrap Elite or LTQ Orbitrap Discovery XL.
Advanced computational modeling and protein design efforts are supported by the IBBR High Performance Computing Cluster (HPCC). The IBBR HPCC contains managed installations of major software packages used in structural biology, molecular dynamics, genomics, and systems biology research such as Rosetta, Amber, PyMOL, and TensorFlow. Hardware resources on the IBBR cluster include 520 CPU cores, permitting large-scale parallel computation, and the cluster recently added 50 NVIDIA GPUs which enable cutting-edge research in machine learning and molecular dynamics simulations. Additionally, our investigators have access to the Maryland Advanced Research Computing Cluster (MARCC) BlueCrab HPCC, a joint center shared by the University of Maryland and Johns Hopkins University, and the Deepthought2 HPCC at the University of Maryland, College Park. Investigators at UGA have also developed EvoMut, a platform for structure-guided biotherapeutic design, which identifies residues that can be modified to increase protein stability by reducing susceptibility to oxidation or protease degradation, or increasing of half-life through glycoengineering.
Glycoprotein analytics
Through the “National Glycoscience Resource” program the CCRC provides services and training for the structural characterization of all types of glycoconjugates to the greater biomedical scientific community including universities, minority serving institutions, federal agencies, biotechnology, and pharmaceutical industries. The particular combination of technologies applied for this purpose will depend on the specific structural information desired by the user and will include LC, mass spectrometry, NMR, and many other technologies.
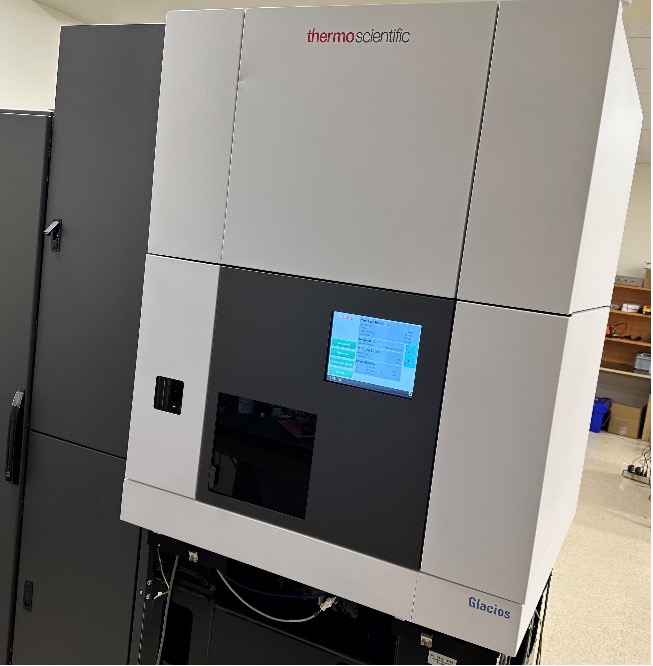

Advanced structural biology tools are supported by the IBBR Keck Center for Structural Biology (Keck Center) and the Maryland Center for Advanced Molecular Analysis (M-CAMA). These tools can be utilized to define key conformations, sites for potential therapeutic interventions, and epitopes important for neutralization of pathogenic infections. Structural information on medically-important proteins using these advanced techniques enables molecular engineering of vaccines that re-focus immune responses to maximize protection and engineering of biologics for optimal activity. Faculty at IBBR have been on the cutting edge of structural biology research for many years, including macromolecular X-ray crystallography and nuclear magnetic resonance (NMR) to determine high resolution structures of small to mid-sized proteins (Keck Center), and, more recently with the opening of M-CAMA, cryogenic electron microscopy (cryoEM), which enables structural characterization of large proteins and complexes that are not amenable to crystallography or NMR (Fig. 6). Taken together, the implementation of all three of these structural methodologies at the CGAB in conjunction with IBBR greatly facilitates efforts in development of vaccines and therapeutics.
Additional QC analysis can entail detection of aggregates using a Zetasizer Nano ZS dynamic light scattering instrument. Confirmation of the identity of proteins produced in the center and monitoring unwanted modifications such as oxidation or deamidation can be accomplished via the mass spectrometry multi-attribute method, which entails peptide mapping and analysis and an LTQ Orbitrap Elite or LTQ Orbitrap Discovery XL.
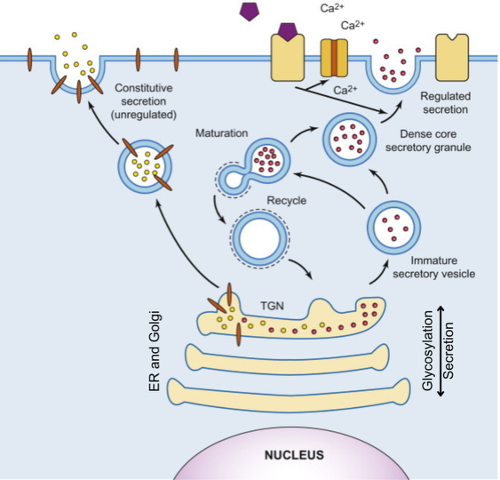
CGAB leverages a comprehensive suite of molecular engineering and cell biology tools to optimize the transit of glycoproteins through the secretory pathway (Fig. 7). Because glycosylation is highly compartmentalized within the endoplasmic reticulum (ER) and Golgi network, the collaboration provides infrastructure to monitor and manipulate protein flux as a means of controlling glycan diversity and density.
Secretion Optimization and Glycoprotein Design: A core capability of the CGAB is the development of platforms to overcome protein retention, a common bottleneck in the production of "glycan-shielded" viral antigens (e.g., SARS-CoV-2, HIV, HCV, and Influenza), and to generate homogeneous glycosylation on key viral and host proteins such as antibodies and serpins. Our researchers utilize AI-guided sequence optimization, signal peptide engineering, and structural biology tools to bypass cellular quality control mechanisms that often lead to low yields or truncated glycosylation. This is supported by: (1) high-throughput secretion assays, (2) live-cell imaging and trafficking analysis, (3) custom cell line engineering, and (4) integrated analytical feedback.
By bridging computational design with structural biology and functional assays, the secretion subgroup provides a closed-loop workflow. We characterize the impact of secretion on glycan occupancy and maturation using the Consortiums's mass spectrometry, ensuring that improved secretion does not come at the expense of glycan fidelity. This integrated approach is essential for the rapid development of next-generation vaccines and biotherapeutics that rely on host machinery for precise post-translational modifications.
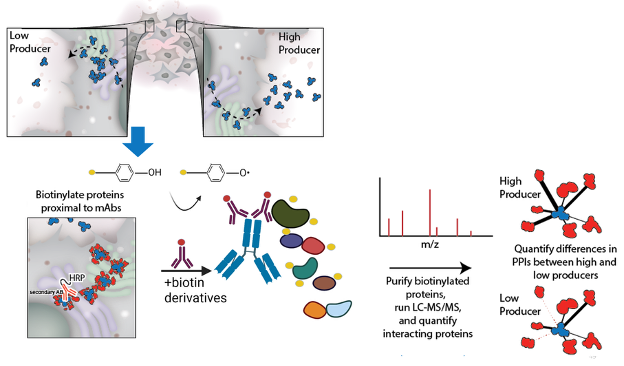
In addition, CGAB researchers have developed a complementary pipeline for enhancing protein secretion yields through the identification of supporting protein-protein interactions, using a suite of proximity proteomics tools, including BioID2, biotinylation by antibody recognition (BAR), and our Fc-specific adaptation FcBAR (Fig. 8). These methods allow us to map, in situ, the host-cell protein interaction networks that enable recombinant protein therapeutics (mAbs, Fc-fusions, secreted enzymes, etc.) to fold, assemble, undergo PTMs, and traffic through the ER/Golgi for secretion. BAR and fcBAR, in particular, use antibody-guided HRP/tyramide labeling in fixed, permeabilized cells to biotinylate proteins proximal to the therapeutic (or endogenous) bait, followed by streptavidin enrichment and LC-MS/MS, capturing transient and compartment-specific secretory interactions. Using this, we have quantified the interactomes of fetuin-B , several SERPIN proteins, our hepatitis C vaccine candidate, multiple mAbs, cytokines, and several other proteins 1-5 .
These approaches allow us to measure PPIs without engineering either the host or the product, enabling clone-to-clone comparisons of what physically helps the molecule get made. Together, these experimental maps feed into our informatics pipeline that integrates proximity MS with transcriptomics and secretory-pathway context to prioritize genetic modifications in production CHO cells to relieve bottlenecks and nominate targeted edits that improve yields of difficult-to-express proteins.
References
- 1. Mina Ying Min Wu, Frances Rocamora, Mojtaba Samoudi, Caressa M. Robinson, Chih-Chung Kuo, Nuša Pristovšek, Lise Marie Grav, Helene Faustrup Kildegaard, Gyun Min Lee, Alexandre Rosa Campos, Nathan E. Lewis. Improving recombinant antibody production using FcBAR: An in situ approach to detect and amplify protein-protein interactions. Metabolic Engineering. Volume 92, 2025, Pages 174-184, ISSN 1096-7176. https://doi.org/10.1016/j.ymben.2025.07.006 . (ScienceDirect) .
- 2. Samoudi M, Kuo C, Robinson CM, et al. In situ detection of protein interactions for recombinant therapeutic enzymes. Biotechnology and Bioengineering. 2021;118:890–904. https://doi.org/10.1002/bit.27621 .
- 3. Helen O. Masson, Mojtaba Samoudi, Caressa M. Robinson, Chih-Chung Kuo, Linus Weiss, Km Shams Ud Doha, Alex Campos, Vijay Tejwani, Hussain Dahodwala, Patrice Menard, Bjorn G. Voldborg, Bradley Robasky, Susan T. Sharfstein, Nathan E. Lewis. Inferring secretory and metabolic pathway activity from omic data with secCellFie. Metabolic Engineering. Volume 81, 2024, Pages 273-285, ISSN 1096-7176. https://doi.org/10.1016/j.ymben.2023.12.006 . (ScienceDirect) .
- 4. M. Y. M. Wu, F. Rocamora, C. M. Robinson, et al. “Enhanced Production of HCV E1E2 Subunit Vaccine Candidates via Protein-Protein Interaction Identification in Glycoengineered CHO Cells.” Biotechnology Journal 20, no. 9 (2025): 20, e70112. https://doi.org/10.1002/biot.70112 .
- 5. Seunghyeon Shin, Frances Rocamora, Nathan E. Lewis. Systematic Mapping of Protein Interactions Underlying IL-2 Secretion in Human T Cells. bioRxiv 2025.09.09.675165. https://doi.org/10.1101/2025.09.09.675165 .